The Bioinformatics Group is actively engaged in developing innovative software tools to advance research and analysis in the field. Explore our projects and find detailed descriptions of our software and usage instructions on group’s GitHub page
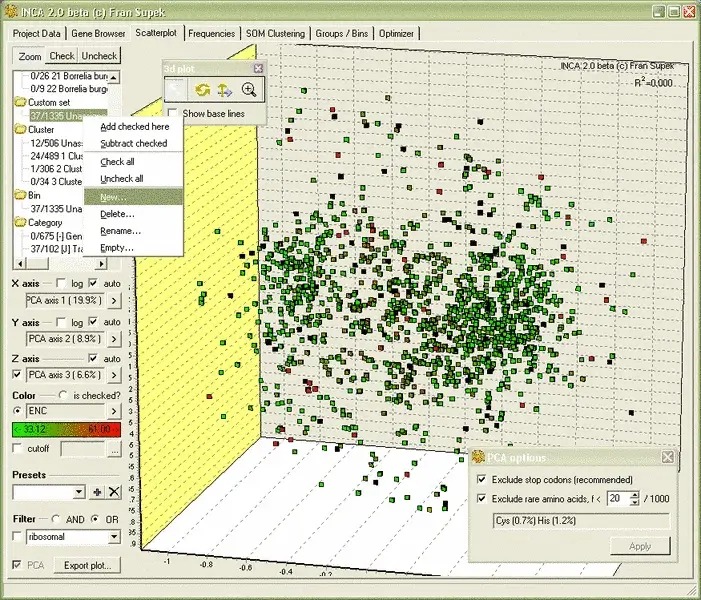
INCA is a powerful tool for analyzing synonymous codon usage in whole genomes. It calculates codon frequencies, indices like codon bias, Nc, and CAI, and provides interactive graphical displays to visualize trends. INCA also includes a self-organizing map (SOM) algorithm for clustering genes based on codon preferences.

coRdon is an R package designed for analyzing and visualizing codon usage in DNA sequences. It calculates various measures of codon usage bias and predictors of gene expressivity. The package performs gene set enrichment analysis for both annotated (KEGG/COG) and unannotated DNA sequences. Additionally, it offers multiple methods for visualizing codon usage patterns and enrichment analysis results.
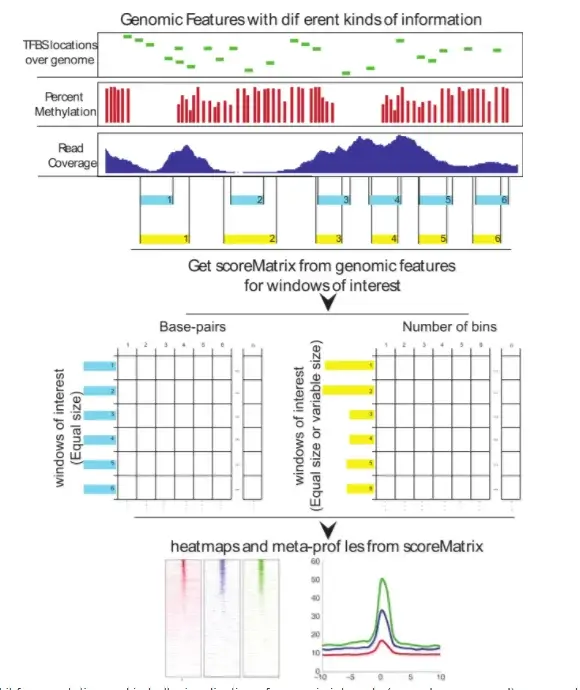
Biological insights can be obtained through computational integration of genomics data sets consisting of diverse types of information. The integration is often hampered by a large variety of existing file formats, often containing similar information, and the necessity to use complicated tools to achieve the desired results. We have built an R package, genomation, to expedite the extraction of biological information from high throughput data. The package works with a variety of genomic interval file types and enables easy summarization and annotation of high throughput data sets with given genomic annotations
Previous projects which are no longer maintained
INCA is a powerful tool for analyzing synonymous codon usage in whole genomes. It calculates codon frequencies, indices like codon bias, Nc, and CAI, and provides interactive graphical displays to visualize trends. INCA also includes a self-organizing map (SOM) algorithm for clustering genes based on codon preferences.
coRdon is an R package designed for analyzing and visualizing codon usage in DNA sequences. It calculates various measures of codon usage bias and predictors of gene expressivity. The package performs gene set enrichment analysis for both annotated (KEGG/COG) and unannotated DNA sequences. Additionally, it offers multiple methods for visualizing codon usage patterns and enrichment analysis results.

While studying molecular biology at the University of Zagreb he developed a strong interest in applying computational methods and approaches to biosciences. For his master thesis he worked on a problem in X-ray crystallography and developed a program for absorption correction of scattered X-ray data. He obtained his PhD in Bioinformatics/Biochemistry, still pursuing his structural biology interests, working on a computational prediction of structural and physicochemical properties of DNA. He spent 10 years as a research fellow at the International Center for Genetic Engineering and Biotechnology in Trieste, Italy, where he also worked on computational prediction and classification of protein domains using machine learning approaches.
In 2002 he established a computational biology group at the Zagreb University, where he moved permanently in 2006 with the EMBO Young Investigators Programme installation grant, and directed the scientific interests towards the newly emerging field of genomics. His bioinformatics group develops computational tools and uses machine learning techniques to tackle open questions in developmental genomics and metagenomics.
In 2011 he became full professor at the Faculty of Science at Zagreb University. He had several international appointments, including a 4-year adjunct professorship at the University of Oslo, Norway and a two-year adjunct professorship at the University of Skövde, Sweden. From 2008 to 2012 he served as the head of the Division of Biology, Faculty of Science at University of Zagreb and was responsible for managing a division of ~150 staff.
He is involved in teaching four graduate-level courses: Bioinformatics, Algorithms and programming, Statistics and machine learning and Computational genomics. During his time as a group leader more than 30 doctoral and master students graduated under his supervision. Some of his graduates continued with PhD and postdoctoral training at prestigious universities around the world, including LMB Cambridge, UK; EMBL, Heidelberg, Germany; RIKEN, Japan and ETH, Zurich, Switzerland.
His scientific track-record includes more than 60 publications in high-level journals and talks in many renowned institutions throughout the world. His research topics are: developmental and differentiation genomics, metagenomics, population genomics and glycomics, origins of multicellularity, epigenomics of cancer, development of computational methods and application of machine learning in genomics and molecular biology.
He is the reviewer in a number of scientific journals, as well as national and international funding bodies and programmes, such as ESF Programme, EC FP7, H2020, and Horizon Europe Programmes, Estonian National Funding Agency, Hungarian National Funding Agency, Flanders Research Foundation, The Netherlands ZonMw Vidi Programme, Croatian National Funding Agencies. He reviewed program applications for Croatian pre-accession and accession structural funds.
Prof. Vlahoviček is a Fellow of the Academia Europaea, and a member of several professional societies, at the national and international level. Professor Vlahoviček is a strong proponent of science reforms in Croatia and evidence-based policy making. He served in the steering committee of Croatia’s highly successful science funding body, the Unity through Knowledge Fund (UKF) and has participated in several strategy-drafting panels at the university and national level. He also served as the Member of the Steering Committee of the Ruđer Bošković Institute, a leading Croatian research entity in natural sciences, with ~900 staff.